 
coli isolates, including dog and human isolates from the same environment. 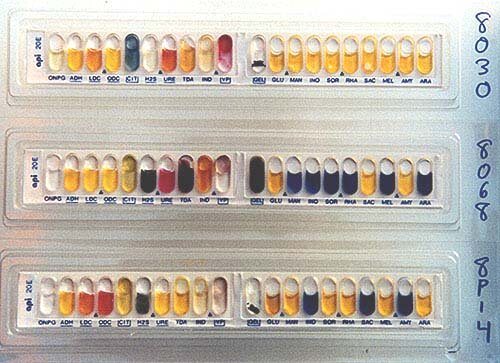
FTIR identified 15 clusters of closely related E. The highest prevalence of ESC-R Enterobacterales faecal carriage was identified in staff working at dog shelters (78.9%), followed by dogs from households (44.11%), dog owners (43.7%), and dogs from shelters (27%). 
Clonal relatedness of canine and human Escherichia coli isolates was established using Fourier Transform Infrared Spectroscopy (FTIR), followed by Illumina WGS of selected isolates. Faecal samples (n = 263) collected from healthy dogs (n = 102), their owners (n = 32), as well as dogs (n = 110) and staff (n = 19) from dog shelters, were screened for ESC-R carriage. This study aimed to determine the prevalence, genetic background, and potential for interspecies transmission of these bacteria between dogs and humans within the same household (HH) or shelter environment in Romania. However, there is no significant difference in the ability of the API 20E to correctly identify "challenge" type organisms (229 of 291) versus routine hospital isolates (71 of 81) (P greater than 0.05), but the system is not as accurate as the conventional biochemical method of identification.įaecal carriage of extended-spectrum cephalosporin-resistant (ESC-R) Enterobacterales in healthy pets is a concerning issue. This evaluation concluded that the accuracy of the identification of Enterobacteriaceae strains at 24 h (78.7%) may be significantly lower than that of earlier evaluations. When 81 of these Enterobacteriaceae strains were arranged into a weighted assortment correlating to the frequency with which they might be found in a clinical laboratory, the API 20E correctly identified 71 (87.7%) at 24 h and 78 (96.3%) at 48 h. The API 20E misidentified eight (2.7%) strains these strains were not limited to any particular genus. At 48 h, 95.2% were correctly identified by using additional biochemical tests as recommended by the manufacturer. At 24 h, the API 20E correctly identified by genus and species 229 of 291 (78.7%) of the strains, using Salmonella and Shigella serotyping where indicated. Because the accuracy of this system compared with conventional biochemical tests has not been determined in many years, we evaluated the API 20E linear strip by using 291 typical and atypical strains of the family Enterobacteriaceae taken from a culture collection. The API 20E bacterial identification system has been used for 19 years, often as the standard with which other identification systems are compared. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
